# Run configuration
base_run_tag = "fixed_term"
experiment_count = 10
max_subjects = 0
# Data parameters
base_data_tag = "HealeyKahana2014"
data_tag = "HealeyKahana2014"
data_path = "data/HealeyKahana2014.h5"
figure_dir = "results/figures"
figure_str = ""
embedding_path = ""#"data/peers-all-mpnet-base-v2.npy"
emotion_feature_path = ""#"data/emotion_features_7col.npy"
feature_column = 6
concat_features = False
trial_query = "data['listtype'] == -1"
target_directory = "results/"
# algorithm selection
model_name = "WeirdCMRNoStop"
make_factory_path = "jaxcmr.models.cmr.make_factory"
component_paths = {
"mfc_create_fn": "jaxcmr.components.linear_memory.init_mfc",
"mcf_create_fn": "jaxcmr.components.linear_memory.init_mcf",
"context_create_fn": "jaxcmr.components.context.init",
"termination_policy_create_fn": "jaxcmr.components.termination.NoStopTermination",
}
sim_alg_path = "jaxcmr.simulation.simulate_study_free_recall_and_forced_stop"
loss_fn_path = "jaxcmr.loss.transform_sequence_likelihood.ExcludeTerminationLikelihoodFnGenerator"
fit_alg_path = "jaxcmr.fitting.ScipyDE"
parameters = {
"fixed": {
"allow_repeated_recalls": False,
"learn_after_context_update": False,
"modulate_emotion_by_primacy": False,
},
"free": {
"encoding_drift_rate": [2.220446049250313e-16, 0.9999999999999998],
"start_drift_rate": [2.220446049250313e-16, 0.9999999999999998],
"recall_drift_rate": [2.220446049250313e-16, 0.9999999999999998],
"shared_support": [2.220446049250313e-16, 99.9999999999999998],
"item_support": [2.220446049250313e-16, 99.9999999999999998],
"learning_rate": [2.220446049250313e-16, 0.9999999999999998],
"primacy_scale": [2.220446049250313e-16, 99.9999999999999998],
"primacy_decay": [2.220446049250313e-16, 99.9999999999999998],
"choice_sensitivity": [2.220446049250313e-16, 99.9999999999999998],
# "emotion_attention": [2.220446049250313e-16, 9.9999999999999998],
# "emotion_scale": [2.220446049250313e-16, 9.9999999999999998],
# "lpp_scale": [2.220446049250313e-16, 9.9999999999999998],
# "delay_drift_rate": [2.220446049250313e-16, 0.9999999999999998],
},
}
# Parameter sweep
varied_parameter = 'start_drift_rate'
sweep_min = 0.
sweep_max = 1.
# Flow toggles
filter_repeated_recalls = True
handle_elis = False
redo_fits = False
redo_figures = True
redo_sims = False
# hyperparameters
seed = 0
relative_tolerance = 0.001
popsize = 15
num_steps = 1000
cross_rate = 0.9
diff_w = 0.85
best_of = 3
# analysis configuration
comparison_analysis_configs = [
# {"target": "jaxcmr.analyses.cat_spc.plot_cat_spc", "figure_suffix": "cat_spc_negative", "kwargs": {"category_field": "condition", "category_values": [1]}},
# {"target": "jaxcmr.analyses.cat_spc.plot_cat_spc", "figure_suffix": "cat_spc_neutral", "kwargs": {"category_field": "condition", "category_values": [2]}},
# {
# "target": "jaxcmr.analyses.nth_item_recall.plot_conditional_nth_item_recall_curve",
# "kwargs": {"query_study_position": 1},
# },
# {
# "target": "jaxcmr.analyses.nth_item_recall.plot_conditional_nth_item_recall_curve"
# },
# {"target": "jaxcmr.analyses.distcrp.plot_dist_crp"},
# {"target": "jaxcmr.analyses.nth_item_recall.plot_simple_nth_item_recall_curve"},
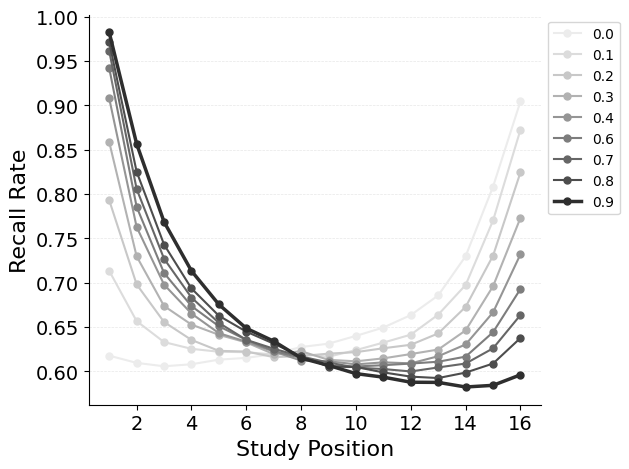
{"target": "jaxcmr.analyses.spc.plot_spc"},
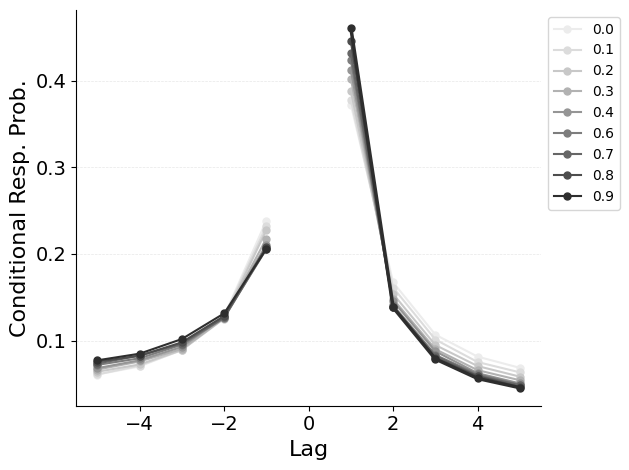
{"target": "jaxcmr.analyses.crp.plot_crp"},
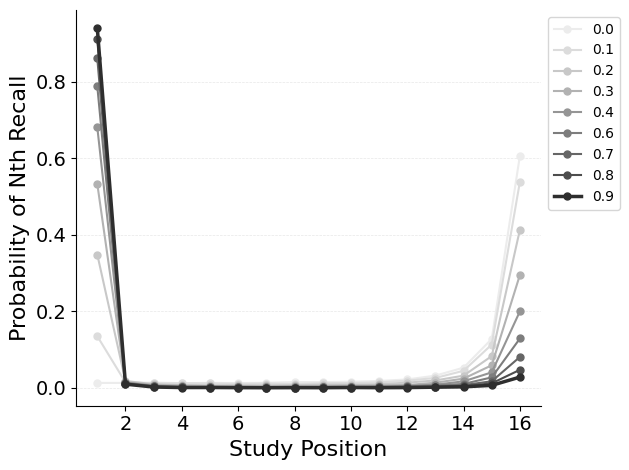
{"target": "jaxcmr.analyses.pnr.plot_pnr"},
# {"target": "jaxcmr.analyses.termination_probability.plot_termination_probability"},
]